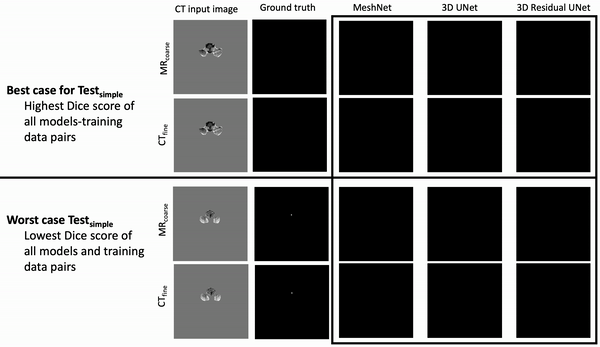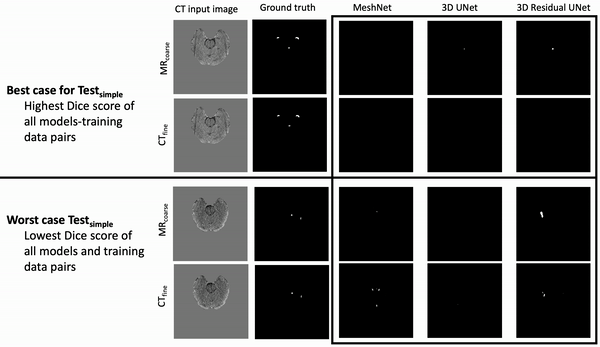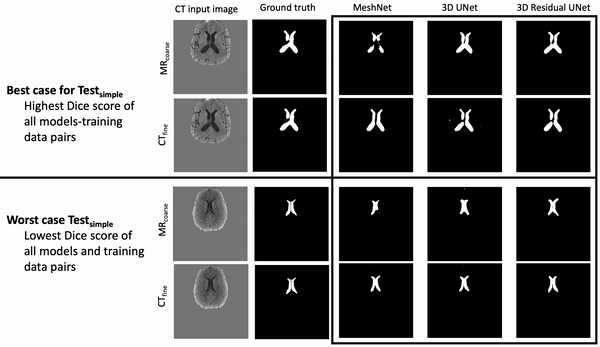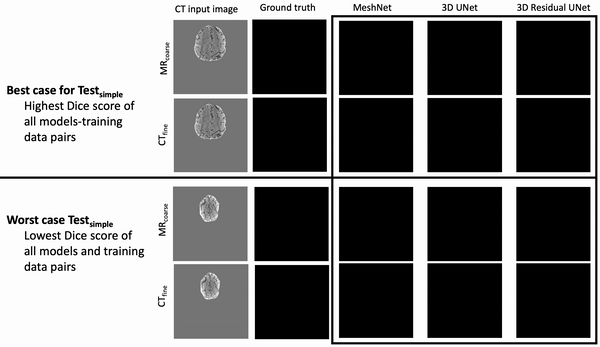
Ventricle Segmentation
Deep Learning Brain and Ventricle Segmentation
TITLE:
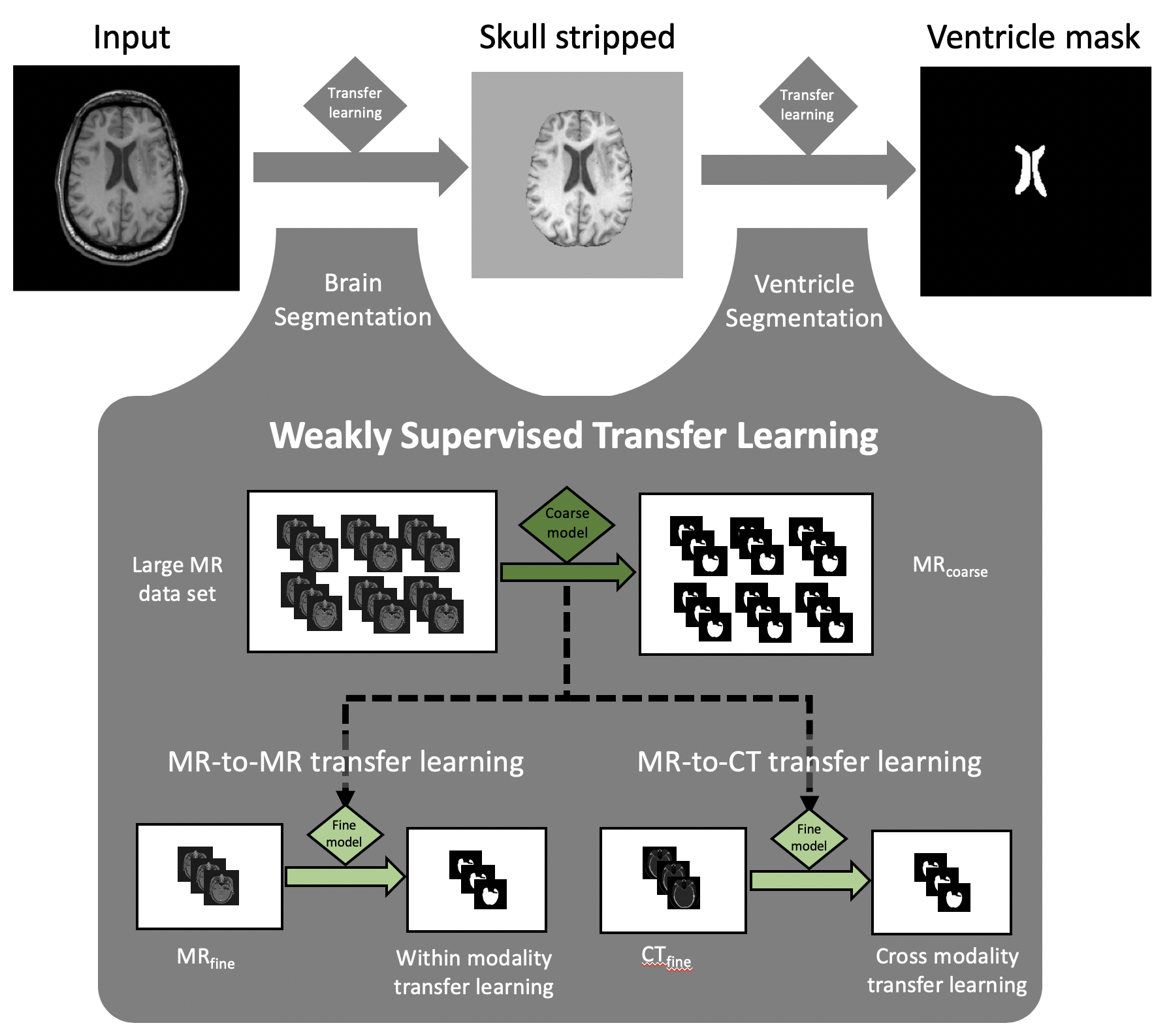
Weakly Supervised Transfer Learning for Multi-Modality Brain and Ventricle Segmentation
Purpose: To quantify performance of various deep learning architectures, models trained with coarse-grained versus fine-grained data, and weakly supervised transfer learning (TL) for segmentation of the brain and ventricles.
Methods: An IRB approved, retrospective study using head MR and CT images of patients was performed. Three datasets consisting of 2193, 162 and 160 images with coarse or fine annotations were used for training, validation and testing of deep learning models. The best model architecture was used investigate the success of TL on segmentation, the influence of training set size for TL accuracy and the effectiveness of cross-modality TL. Dice score (DS) and percent volume difference (PVD) were used to quantify model performance. Two-sided Wilcoxon signed rank with p>0.05 indicated statistical significance.
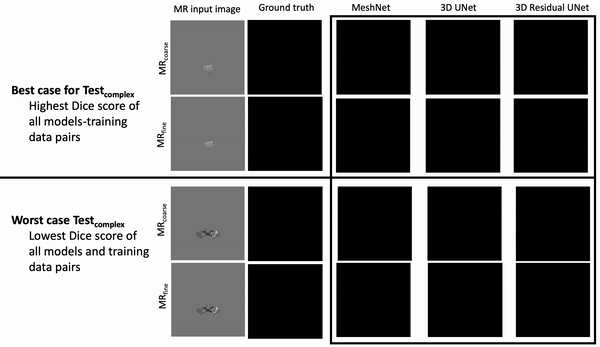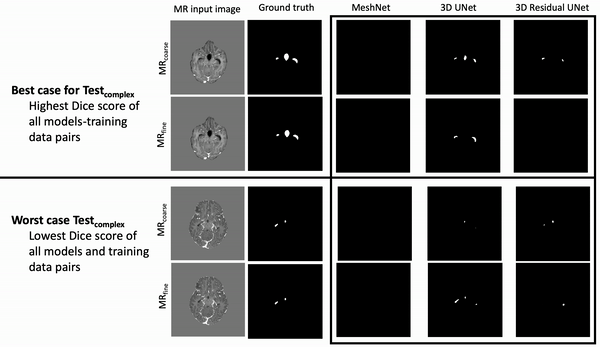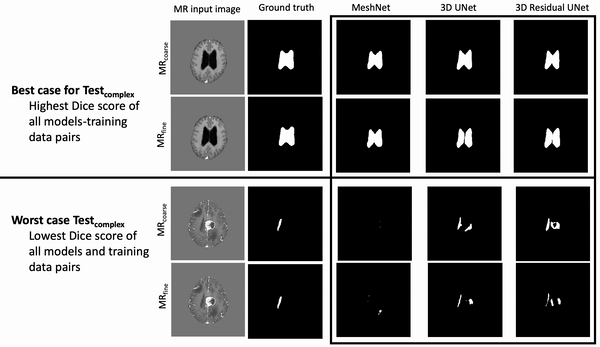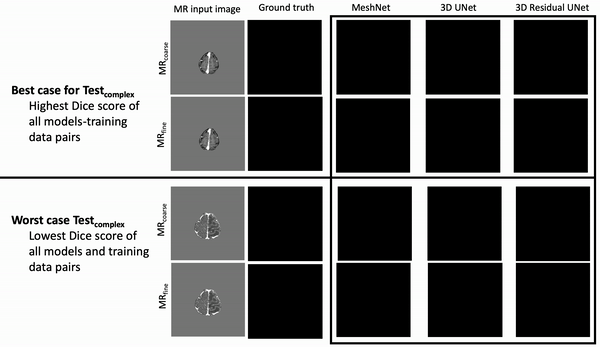
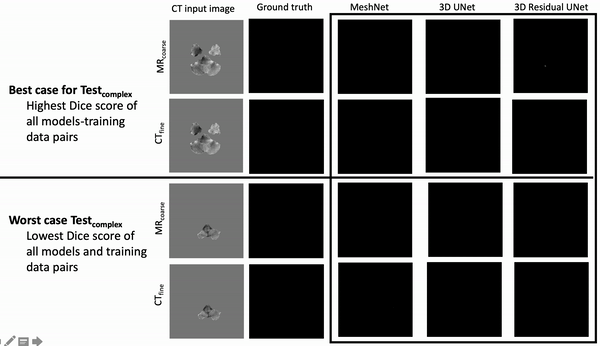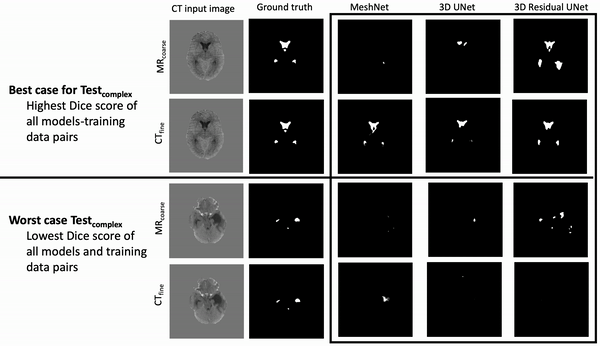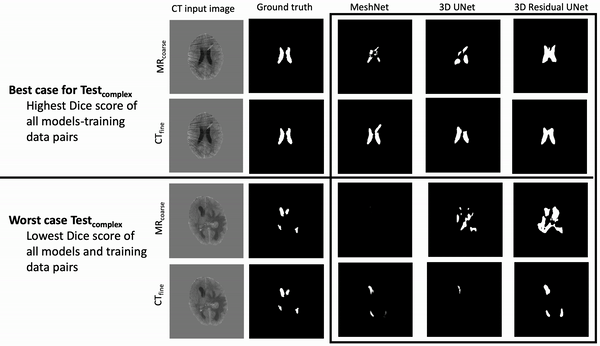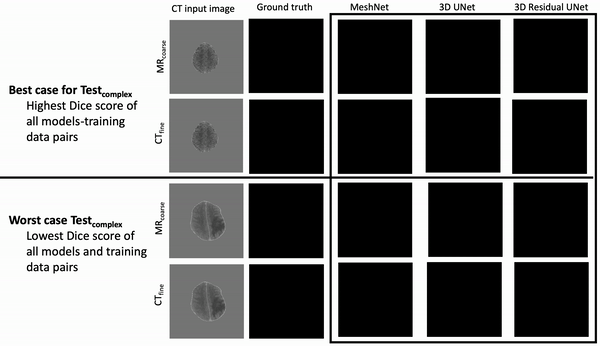
Results: Deeper, wider (more parameters and outputs) models outperformed other architectures for segmentation tasks. Most brain segmentation models were statistically equivalent. Ventricle segmentation models trained with fine-grained data improved DS from 0.75 to 0.82 and PVD from 21.5% to 8.02% over coarse-grained trained models, both results statistically significant. Both DS and PVD improved when using TL over noTL (0.86 vs. 0.83, p < 0.01 and 6.87% vs. 8.02%, p = 0.80, respectively). TL models trained with as few as 20 images showed similar results to models trained with 100 images and vastly outperformed de novo models.
Conclusion: Weakly supervised TL segmentation models utilizing deep, wide networks trained with small volumes of fine-grained data outperform shallow networks trained on large volumes of coarse-grained data and to models without TL.
Python code development & Data for the project:
https://github.com/medomics/MEDomicsLab-ventproject